Proximity RNA labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules | bioRxiv

Enzymatic RNA Biotinylation for Affinity Purification and Identification of RNA–Protein Interactions | ACS Chemical Biology

mRNAs biotinylated within the 5′ cap and protected against decapping: new tools to capture RNA–protein complexes | Philosophical Transactions of the Royal Society B: Biological Sciences

Synthesis of biotin–AMP conjugate for 5′ biotin labeling of RNA through one-step in vitro transcription | Nature Protocols
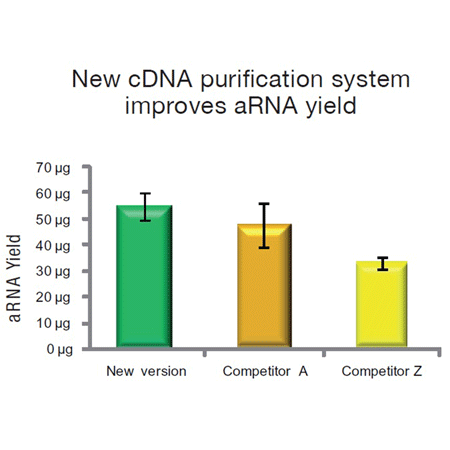
BIOARRAY™ Single-round RNA amplification and biotin labeling system - ENZ-42420 - Enzo Life Sciences
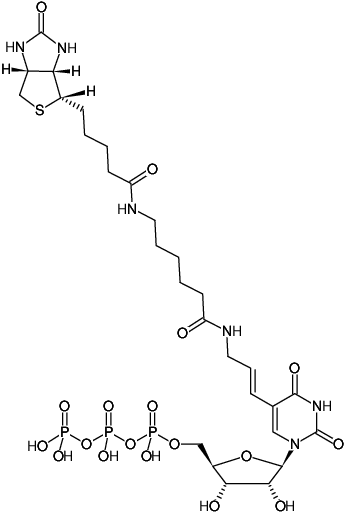

HighYield T7 Biotin11 RNA Labeling Kit (UTP-based), Random RNA Labeling (in vitro Transcription-based) - Jena Bioscience